Call for Open-Source Projects, Scaling of NNPs, and More Labs
request for open-source scientific packages; an opinionated guide to open-source scientific-software development; getting physical scaling correct without physics; two new labs for the classroom

Although we don't have any new features this week, there's a variety of articles, guides, and labs that we're excited to share.
Open-Source Projects We Wish Existed
At Rowan, we work to support scientists in many different areas of chemistry: we benchmark existing models and algorithms, write our own code, and try to build robust and accurate solutions for our users and customers.
As a part of this work, we spend a lot of time working with open-source scientific code. We’ve contributed a bit to open-source software already: Corin maintains pymysym and cctk (among others), Jonathon maintains a scientific cookiecutter for Pixi and steamroll (a package for converting 3D molecular coordinates to RDKit objects), and we released Egret-1, a family of MIT-licensed NNPs, earlier this year. (There are more open-source projects to come!)
Still, we often find ourselves wishing that there were a specific open-source library that we could use for some task or another. After a variety of one-on-one conversations with academics and industry scientists interested in contributing to useful scientific software development, we realized it would be useful to collect these ideas in one place.
Projects we imagine include:
New optimizers designed to work well with neural network potentials,
A package to combine different SCF-convergence strategies,
And an improved way to go from 3D coordinates to "2D representations" (RDKit objects or SMILES strings).
If you are looking for research projects or want to get better at developing scientific software, we think these are projects that will be very useful to the computational chemistry community. We’d be happy to talk with you about whichever of these you find interesting.
How to Make a Great Open-Source Scientific Project
The process of developing new scientific packages can be daunting, especially for newcomers to programming. As avid developers and users of many open-source projects like the projects listed above and community projects like Psi4, the RDKit, and xtb, we’ve had the opportunity to work with many different packages and learn what is and isn’t useful when developing new packages or maintaining existing ones. The best codebases can be a delight to work with and are often tightly focused, well documented, and regularly updated on PyPI or Conda.
Getting your project to this point can be daunting: “How does one maintain dependencies?” “What type of testing should a project have?” “How do you have high code-quality and fast development of new features?” These things are rarely taught, but instead slowly absorbed over years of programming, leading newcomers to have to learn via trial and error. We've put together a guide that we believe will help scientists build great open-source projects the community can easily adopt.
This is an opinionated guide to Python-based scientific projects, although we hope parts of this guide will be applicable to other languages as well. For those starting new projects, the pixi-cookiecutter provides a simple way to start projects that follow these guidelines, making it easy to write and maintain high-quality code. For those working on existing projects, we recommend focusing on the simplest additions first (e.g. using an autoformatter, having tests run when code is merged, and packaging code in a manner that is easy for others to use)—there's no need to change everything immediately. Read our guide covering the eight attributes of good open-source projects here.
Studying Scaling in Electron-Affinity Predictions
Last week, we released a paper showing that OMol25-trained NNPs can accurately predict molecular properties that depend on the difference in energy between different charge and spin states, like reduction potential and gas-phase electron affinity. Across a wide variety of main-group and organometallic systems, we observed that NNPs could be used as a high-throughput way to predict these properties with good accuracy.
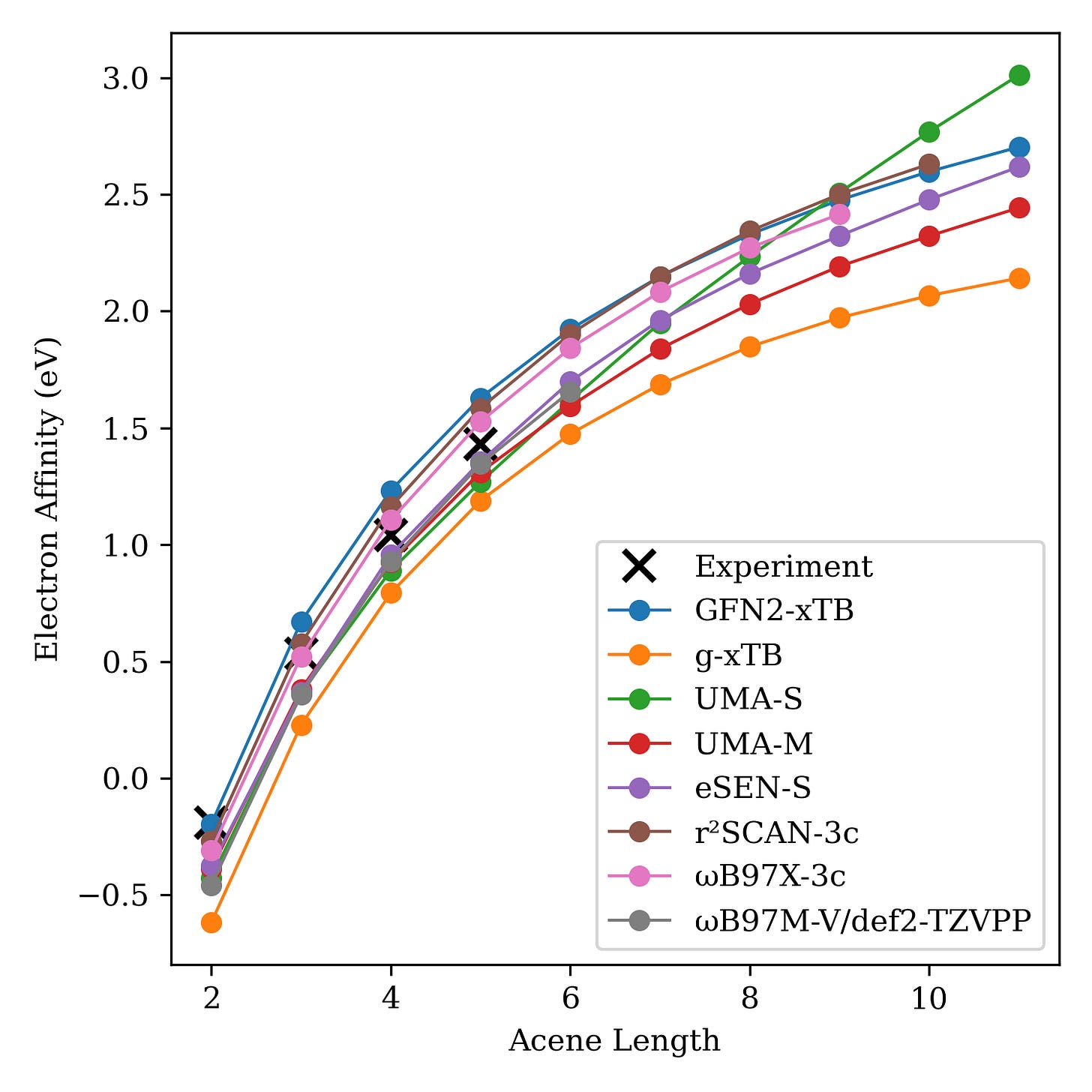
Since these models were trained on so much data, it's possible that the observed good performance is simply a form of "memorization" and that non-physical behavior will occur as we scale to larger and larger systems. To test this, we examined a series of acenes of increasing length to see if the NNPs could properly capture the electron-affinity trend. The results surprised us and suggest that NNPs are actually learning chemistry beyond just a simple "lookup table" model, although this is just one study. Read the full blog post (with data and code) here.
Trends in Acidity Lab
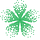
Acidity is a concept often first learned as a child when mixing baking soda and vinegar. It is then introduced formally in introductory chemistry classes, with various labs requiring careful acid–base titrations to measure the pKa of a single molecule. Trends are memorized and repeated, but a fundamental hands-on understanding is rarely possible due to the time required for each titration.
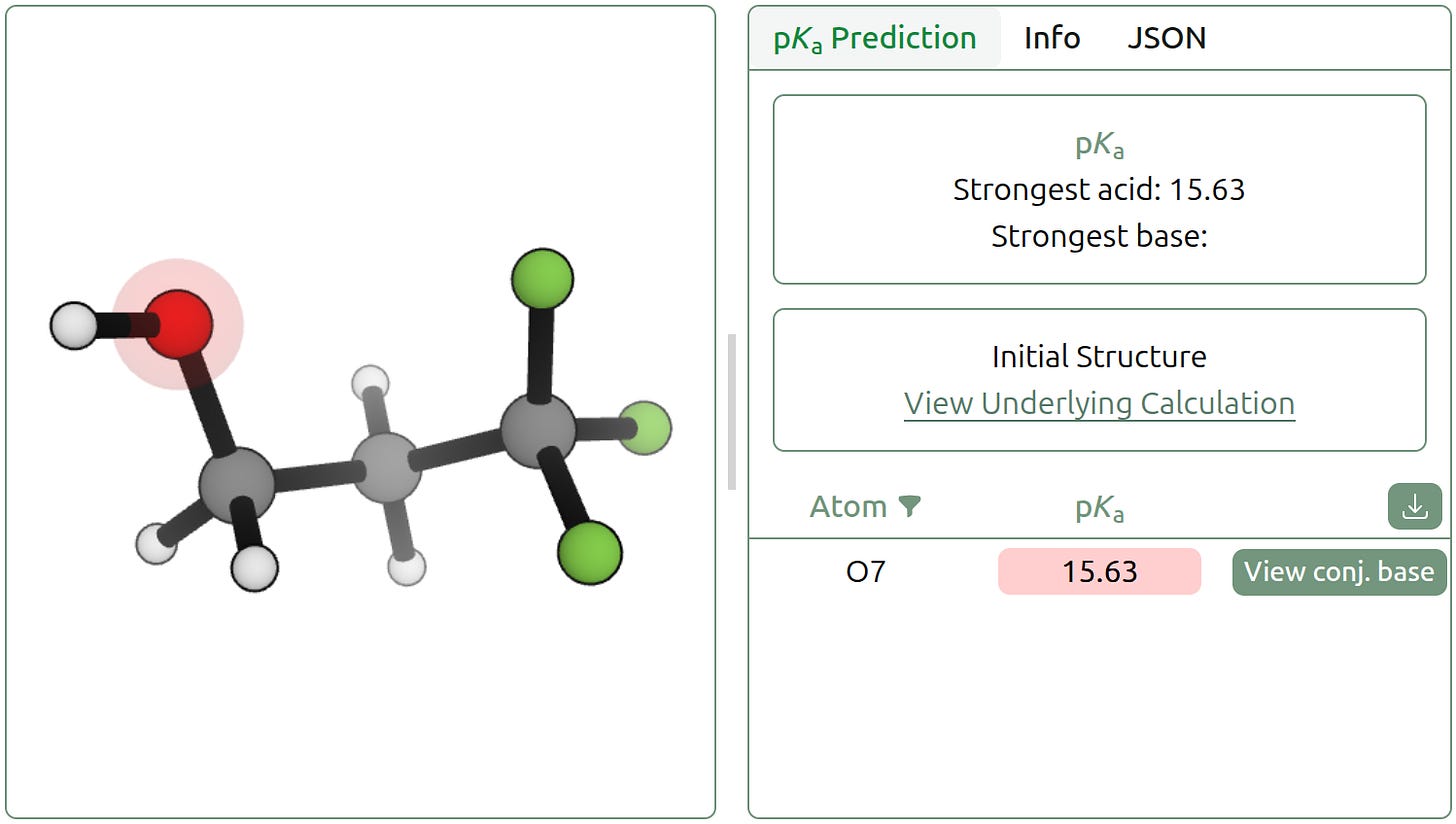
Jay Sahni, a student at Jordan High School in Texas, decided he wanted to help us make the concepts behind these trends more concrete. He wrote a lab about the trends in acidity intended for introductory organic chemistry students. This lab is written to encourage exploration. Students can quickly predict the pKa of many different molecules using the Rowan platform, allowing them to independently experiment with a variety of different electron-withdrawing and electron-donating substituents to determine their effects on acidity, and gain their own understanding of various trends.
Bond-Dissociation Energy Lab
While the pKa lab above is concerned with heterolytic cleavage (where the electrons both go to the same atom) and the resulting solvent stabilization of the ions, our summer intern Sawyer VanZanten wrote a lab about homolytic bond cleavage (where the electrons split between the atoms), which is often used as an indicator of bond strength. Bond strengths can help predict the regioselectivity of C–H functionalization reactions or potential sites of metabolism for drugs, and the concept of bond strength is taught in introductory chemistry classes to help rationalize why certain reactions can happen.
This lab introduces students to the concept of bond-dissociation energy (BDE) and helps students rationalize the differences between the strengths of various bonds via hands-on predictions of bond-dissociation energies using the Rowan platform.




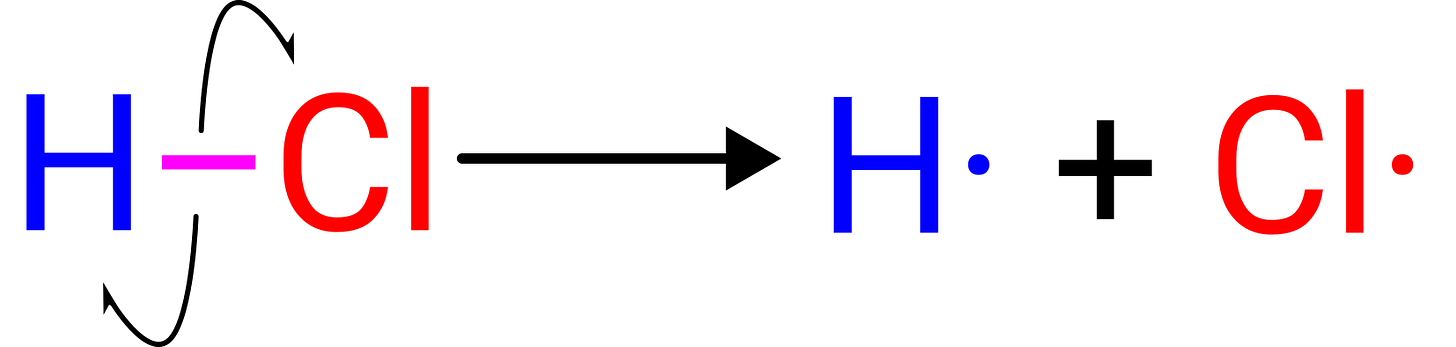
I had to double take, I think the homolytic and heterolytic have gotten mixed up here. (The acid should be heterolytic)