Predicting Permeability for Small Molecules
why permeability matters; different experimental and computational approaches; Rowan’s supported methods; an example script
Today, we’re launching a permeability-prediction workflow on Rowan. Membrane permeability, often abbreviated to just “permeability” in drug-discovery contexts, quantifies how quickly a molecule can get across a given lipid membrane. This matters at numerous stages in the drug-delivery process:
If a drug is taken orally, then it must pass through the epithelial membrane in the intestine to enter the bloodstream.
Once in the bloodstream, the drug must pass through cell membranes in order to reach intracellular targets.
And drugs targeting the central nervous system must also pass through the blood–brain barrier, an additional membrane that protects the brain from potentially harmful xenobiotics.
In the words of the mighty Derek Lowe, “membrane permeability is one of the most basic properties that we look for in a drug.” Accordingly, it’s common for drug-discovery scientists to test permeability extensively in preclinical research. There are a lot of experimental assays that are used: cell lines like Caco-2 and MDCK are often used to measure permeability, as are artificial membranes like PAMPA (for more details, see this excellent analysis by Karel Berka and co-workers). However, all of these experimental assays require a sample of the synthesized compound and cost additional time & money, limiting the potential throughput.
Since permeability is so crucial, it’s become common to predict drug permeability computationally before actually synthesizing or ordering new compounds. While computational models are never as accurate as actually running the experimental assay, they can be run quickly and cost-effectively on thousands or tens of thousands of compounds, allowing scientists to avoid molecules predicted to have bad permeability and spend their limited time and resources only on the most promising candidates.
We’re excited to be launching two complementary approaches to permeability prediction for Rowan subscribers today.
The Machine-Learning Approach: GNN-MTL
Our first permeability-prediction method comes from Philip Ohlsson and co-workers at AstraZeneca, who recently published a paper in ACS Omega showing that multi-task learning could improve the performance of graph neural networks for permeability prediction. The model associated with this paper was trained on large amounts of internal data from AstraZeneca and is open source; we’ve added this model to Rowan.
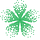
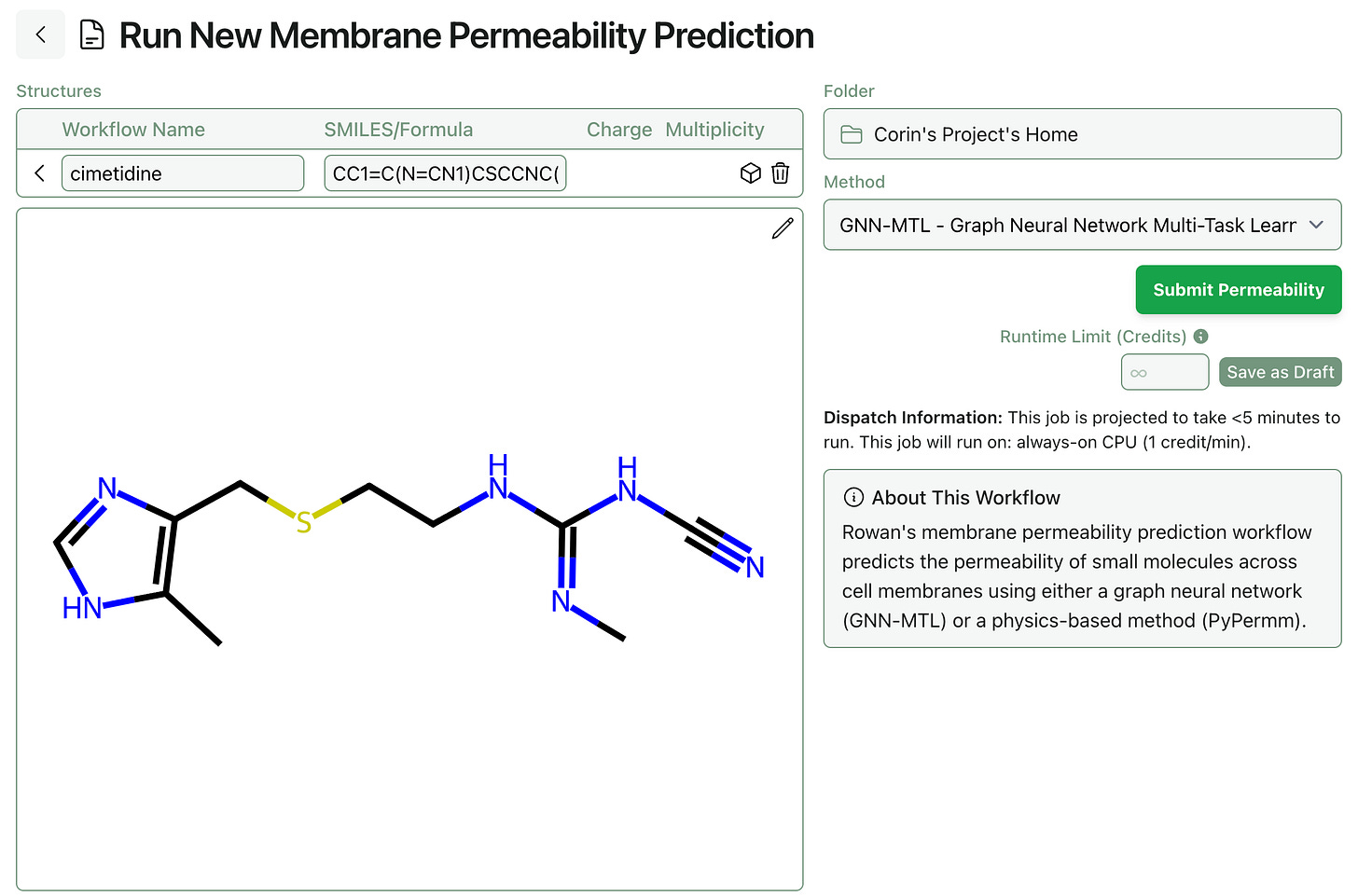
To run a prediction with Ohlsson’s model, simply select the new “Membrane Permeability” workflow and input a 2D structure of the desired molecule.
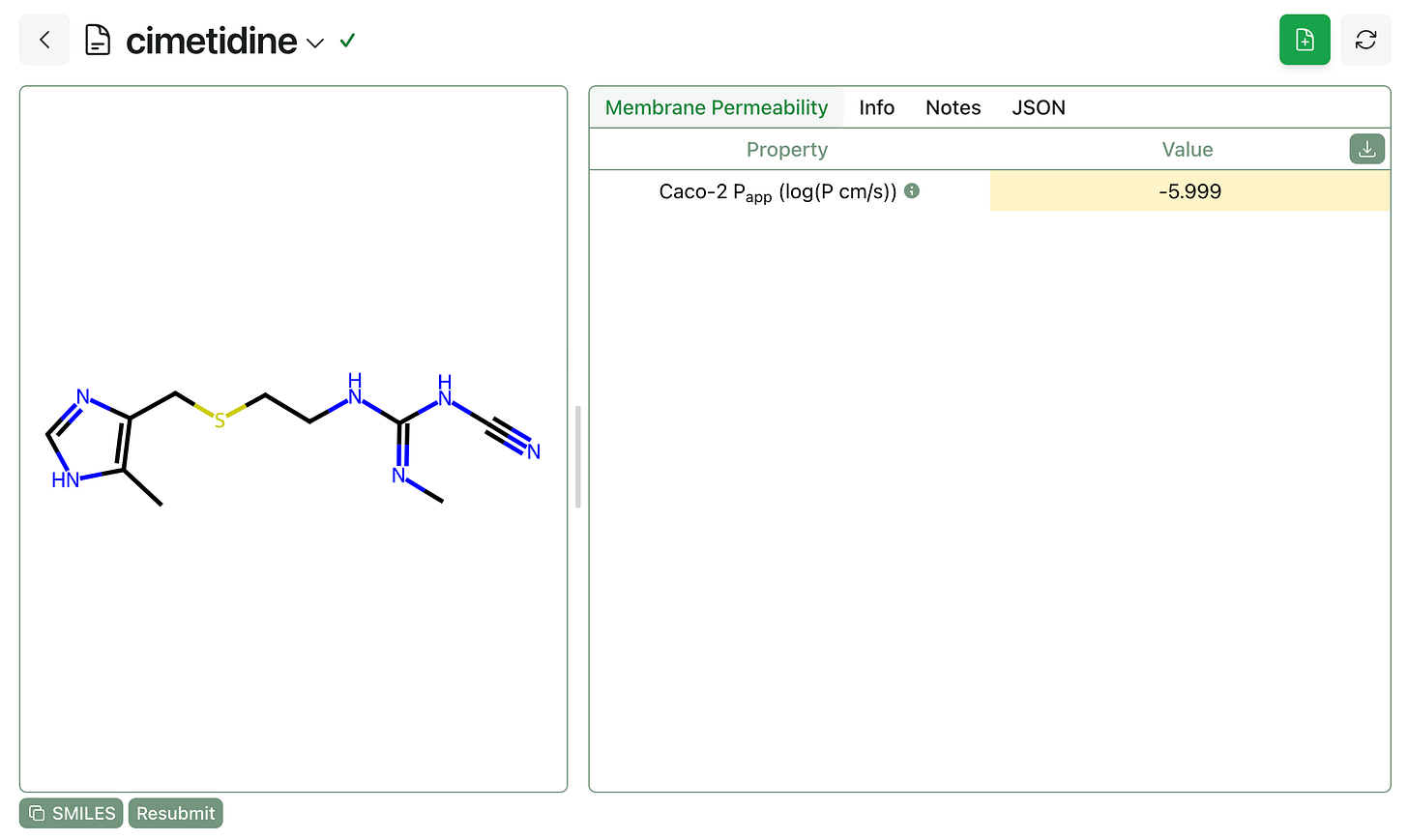
The GNN-MTL model takes a few seconds to run and returns the predicted Caco-2 apparent permeability in cm/s. We’ve color-coded the outputs to help translate the output values into qualitative terms; cimetidine below is predicted to have mediocre permeability.
The Physics-Based Approach: PyPermm
A totally different approach to predicting permeability comes from Andrei Lomize and co-workers, who developed a physics-based method for modeling passive diffusion across membranes. Lomize and co-workers previously released the PerMM web server, which makes it possible to compute free-energy profiles and intrinsic permeability coefficients for arbitrary small molecules. We’ve reimplemented PerMM in Python and released it as PyPermm on GitHub.
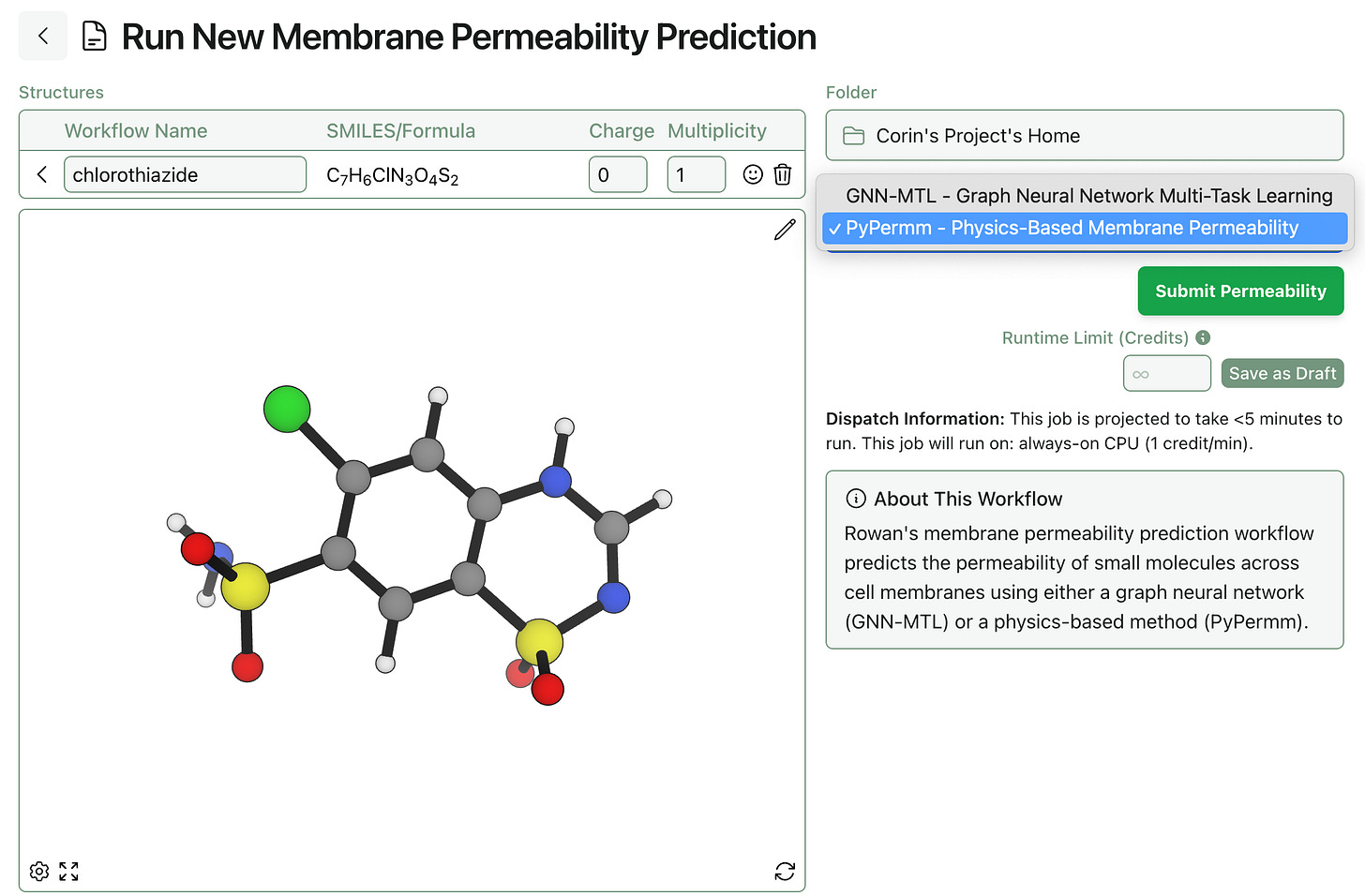
To run PyPermm-based permeability predictions in Rowan, simply select the PyPermm method on the workflow submit screen. PyPermm requires all-atom 3D structures to run, unlike GNN-MTL:
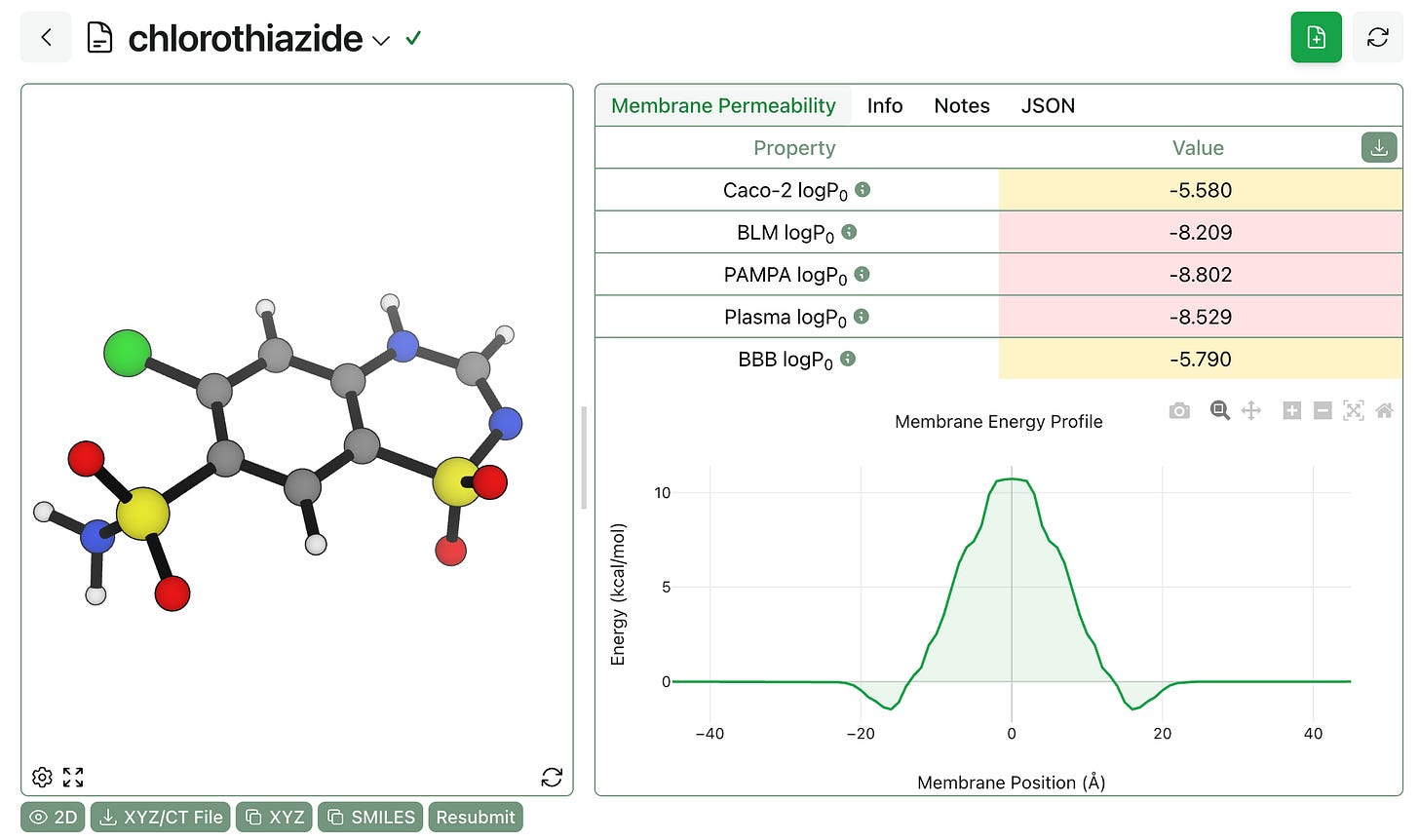
PyPermm returns (1) predicted intrinsic permeability values for a variety of different membrane types and (2) the predicted free-energy profile for membrane transport. Here’s what this looks like for chlorothiazide; as with GNN-MTL, we’ve added colors to indicate qualitative permeability.
Both methods can easily be run through Rowan’s Python API:
from rdkit import Chem
import rowan
smiles = "CC1=C(N=CN1)CSCCNC(=NC)NC#N"
gnn_mtl_result = rowan.submit_membrane_permeability_workflow(
smiles,
name="GNN-MTL permeability",
)
pypermm_result = rowan.submit_membrane_permeability_workflow(
Chem.MolFromSmiles(smiles),
method="pypermm",
name="PyPermm permeability",
)
gnn_mtl_result.wait_for_result().fetch_latest(in_place=True)
pypermm_result.wait_for_result().fetch_latest(in_place=True)
print(f"Caco-2 Papp (GNN-MTL): {gnn_mtl_result.data['caco_2_P_app']}")
print(f"Caco-2 P0 (PyPermm): {pypermm_result.data['caco_2_logP']}")Rowan’s permeability-prediction workflow is now live for all subscribing users, and we look forward to hearing feedback on how these models perform for individual projects. As new permeability-prediction models are released, we anticipate adding new methods to Rowan, so please let us know if you see a paper that you’re interested in trying out!